PASSC is a command line program which compares a protein sequence to another protein sequence-structure data obtained from SHEBA, or to a protein sequence-structure database. It gives the pair-wise alignment and the statistics for multiple alignment. The input and output files are:
1. (*.seq): a query sequence file in FASTA format. It begins with a single line description, starting with a greater-than (">") symbol in the first column, and followed by lines of sequence data in a separate line. Sequences are expected to be represented in the standard IUB/IUPAC amino acid codes. Here is an example:

2. (.psc): a subject sequence file in PASSC format. In addition to the amino acid sequence in FASTA format, the subject sequence for the PASSC program has secondary structure and polarity information which are derived from the (*.env) files created by SHEBA. The PASSC format is like this:

It begins with a single line description, starting with a greater-than (">") symbol in the first column, and followed by lines of amino acids in CAPITAL case. The secondary structure of each residue is represented in 1 of the 4 characters in smaller case: c for coil, h for helix, s for sheet, and t for turn. Numbers 0, 1, 2, or 3 are meant to be the polarity exposure of each amino acid 0% ~ 25%, 25% ~50%, 50% ~ 75%, or 75% ~ 100%, respectively. The command mkpsc in the io option or the stand-alone function mkpsc can be used to generate the PASSC format file.
3. (.lib): the list of (.psc) files:
For one-to-many alignment, PASSC needs the subject library, which has concatenated (*.psc) files like below. The library file needs at least 20 proteins in order to get meaningful statistics.

4. The Hk2 and Tk2 matrices in binary format:
The Hk2 score matrix is in the PAH.score and Tk2 score matrix is in the SEC.score. Those two matrices in binary format are required for PASSC and are created by HscoreTbin and SscoreTbin. To generate, please see the example page.
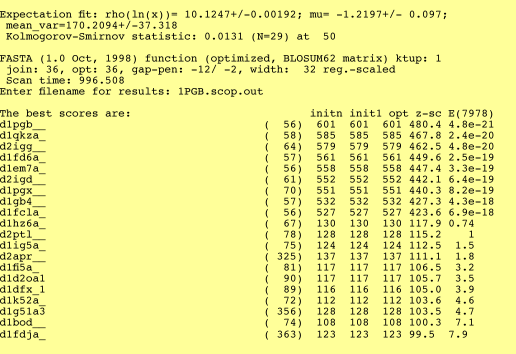
5. Alignment, histograms, and statistics report output:
PASSC has 3 different kinds of output: pair-wise alignment, histogram, and statistics report. In this latest version (2.2), the secondary structure and polarity information of the subject sequence were added in the alignment output. The alignment is based on the Smith-Waterman algorithm and modified with Hk2 and Tk2 matrices as shown below.

In the histogram, the expected number of sequences is plotted using an "*". The z-score is in the first column, the second column is the number of proteins and the third column is the E() value. 
The statistical routines assume that the library contains a large sample of unrelated sequences. If this is not the case, then the expectation values are meaningless. Likewise, if there are fewer than 20 sequences in the library, the statistical calculations are not done. The random average score is displayed in the column initn. For protein searches, library sequences with E() values < 0.01 for searches of a 10,000 entry protein database are almost always homologous. Frequently sequences with E()-values from 1 -10 are related as well. Remember, however, that these E() values also reflect differences between the amino acid composition of the query sequence and that of the "average" library sequence.