LSHEBA (Local Structual Homology by Environment-Based Alignment)
Examples (Cont'd)
IV. Other Output:
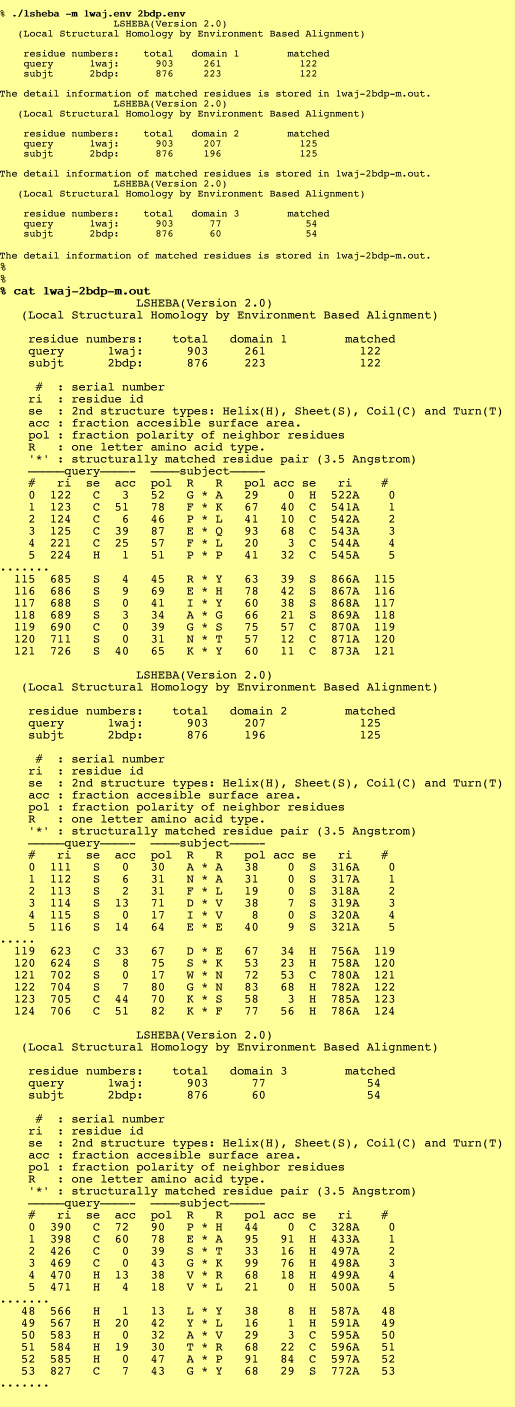
1. Detail information of matched residues:
This output lists the residue id, secondary structure code, accessibility, and polarity of the matched residues.
%./lsheba -m (query.env/.pdb) (subject.env/.pdb)
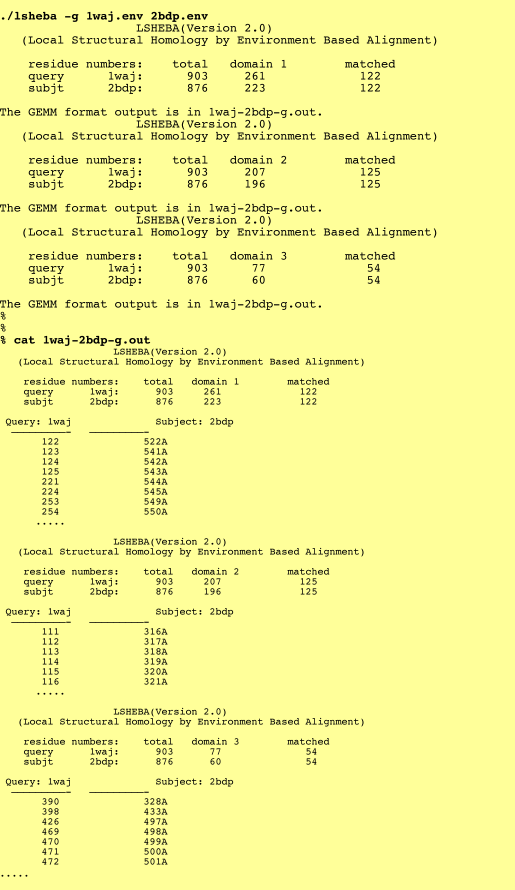
2. GEMM format output:
This format can be parsed to GEMM and it is stored in a file. Default output will be on the screen.
% ./lsheba -g (query.env/.pdb) (subject.env/.pdb)
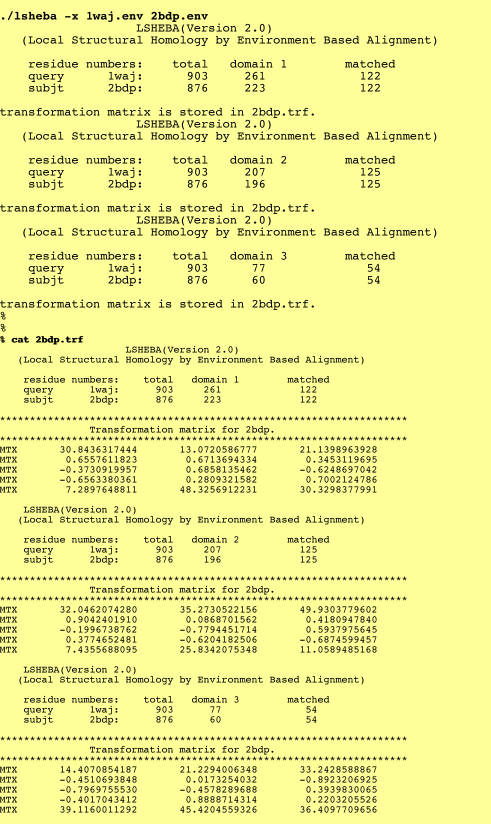
3. Transformation matrix:
The transformation matrix of each domain will be saved in one file and default output is on the screen.
%./lsheba -x (query.env/.pdb) (subject.env/.pdb)