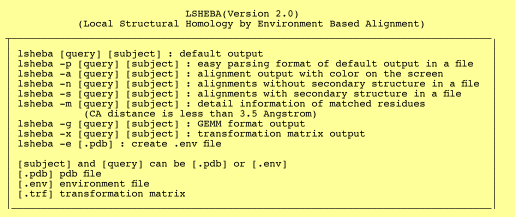
LSHEBA is a command line program that finds the local alignments of two proteins. It has the following options and the help manual can be found by typing lsheba. It gives the statistics as well as the alignment, matched residues and transformation matrix. To learn more, please click on each of the options below and the example page will be displayed.
The input and output files are very similar to SHEBA program:
1. (.pdb): a flat PDB (ascii) format file. If the PDB file name ends with (.ent), please re-name it as (.pdb) so that LSHEBA can recognize. If the PDB file comes from an NMR experiment, there might be more than one model and LSHEBA only processes the first one.
2. (.env): a PDB-like format with only alpha carbon coordinates and enviornment values. Here is the example:

3. (.trf): an ascii file that contains a transformation matrix. One can obtain this transformation matrix by giving a LSHEBA command with "-x" option. This matrix transforms the second pdb coordinates toward the first one.